DCOR access
The deformability cytometry open repository (DCOR) allows you to upload and access RT-DC datasets online (internet connection required). The advantage is that you can access parts of the dataset (e.g. just two features) without downloading the entire data file (which includes image, contour, and traces information).
Public data
When you would previously download an entire dataset and do
import dclab
ds = dclab.new_dataset("/path/to/Downloads/calibration_beads.rtdc")
you can now skip the download and use the identifier (id) of a DCOR resource like so:
import dclab
ds = dclab.new_dataset("fb719fb2-bd9f-817a-7d70-f4002af916f0")
To determine the DCOR resource id, go to https://dcor.mpl.mpg.de, find the resource you are interested in, scroll down to the bottom, and copy the value from the id (not package id or revision id) field in (Additional Information). The DCOR format is documented in DCOR (online) format.
Private data
If you want to access private data, you need to pass a personal API Token.
import dclab
ds = dclab.new_dataset("fb719fb2-bd9f-817a-7d70-f4002af916f0",
api_key="XXXX-YYYY-ZZZZ")
Alternatively, you can also set an API Token globally using
import dclab
from dclab.rtdc_dataset.fmt_dcor.api import APIHandler
APIHandler.add_api_key("XXXX-YYYY-ZZZZ")
ds = dclab.new_dataset("fb719fb2-bd9f-817a-7d70-f4002af916f0")
Managing API Tokens
You can manage your API Tokens on your profile page when logged in at https://dcor.mpl.mpg.de.
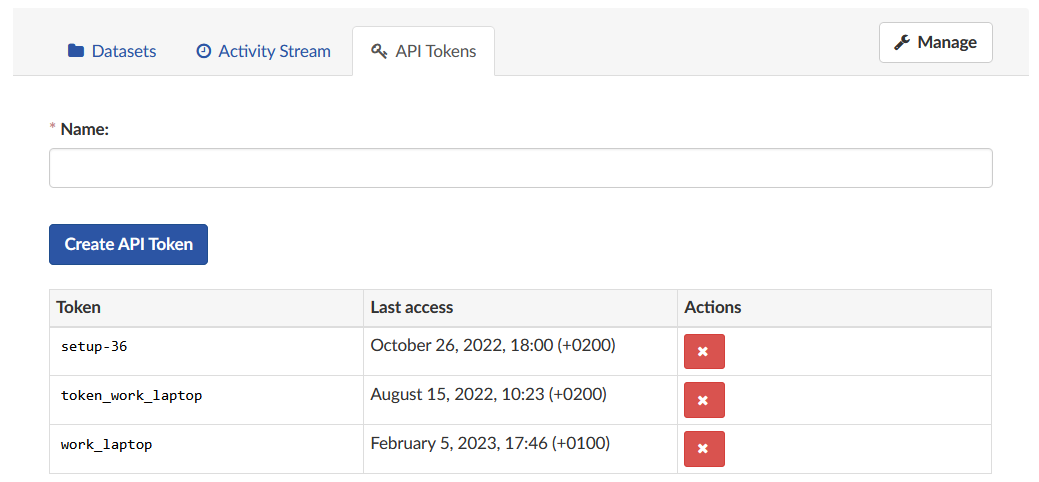
Fig. 5 Managing API Tokens on DCOR.
Deleting a token:
Click on the tab “API Tokens” to view all currently existing tokens and the date they were last accessed. By clicking on the red “X” you can delete a token. It cannot be restored, so be careful when deleting tokens!
Creating a new token:
To create a new token, insert a name in the field at the top and click “Create API Token”. The newly generated token will be shown at the top of the page. Make sure you copy it directly, because you will not be able to recall it again!
Accessing data on a different DCOR instance
To access data on a different DCOR instance, you have to pass the respective
URL when opening the dataset via the keyword host. The procedure to
retrieve the DCOR resource id is the same as for the default DCOR.
import dclab
ds = dclab.new_dataset("fb719fb2-bd9f-817a-7d70-f4002af916f0",
host="dcor-dev.mpl.mpg.de")
Bypassing DCOR and using S3 directly
The DCOR format connects to the dcserv API on on the DCOR server side. Internally, DCOR uses an S3-compatible object store to manage all resources. In some scenarios you might want to bypass this API and access individual DCOR resources directly.
Advantages:
potentially faster access to HDF5 data using the S3 format or other software, since the
dcservwrapper is bypassedyou don’t have to depend on dclab in your code
Disadvantages:
no direct access to private resources: You either need to use the
dcservAPI to obtain a presigned S3 URL (which also has an expiry date) or you need to own S3 credentials for the object store.no direct access to features from the condensed file: DCOR automatically computes a condensed file upon upload. This file contains only (but more) scalar features. The
dcservAPI transparently combines features from the original and the condensed file.
Resources are stored in the following pattern by DCOR:
https://{endpoint_domain}/{instance-specific-prefix}{circle-id}/resource/{resource-id}
For instance, the calibration beads dataset, has this S3 URL:
https://objectstore.hpccloud.mpcdf.mpg.de/circle-5a7a053d-55fb-4f99-960c-f478d0bd418f/resource/fb7/19f/b2-bd9f-817a-7d70-f4002af916f0
You can access condensed resources by replacing resource with condensed in the above URL:
https://objectstore.hpccloud.mpcdf.mpg.de/circle-5a7a053d-55fb-4f99-960c-f478d0bd418f/condensed/fb7/19f/b2-bd9f-817a-7d70-f4002af916f0